Projects
International Cancer Genome Consortium (ICGC) and PCAWG
Laboratory for Cancer Genomics
Laboratory for Medical Science Mathematics
The ICGC was established in 2007, and concluded its mission to define the genomes of 25,000 primary untreated cancers (the 25K Initiative) in May 2018. The ICGC solved numerous data governance, ethical and logistical challenges to make global genomic data sharing for cancer a reality, providing the international community with comprehensive genomic data for many cancer types. As the second ICGC initiative, the ICGC launched a “Pan-Cancer” Whole Genome project (PCAWG) in 2014, in which WGS data together with RNA-Seq of 2834 donors were analyzed in uniform pipelines within the same computational environment and cloud computing. RIKEN and IMUT (Institute of Medical Sciences, The University of Tokyo) are contributing to this project as a member of a technical working group arranging ten cloud data centers and PI/researchers in working groups for driver gene, mutational signature, germline, immuno-genomics, and mitochondrial genomics. RIKEN provided WGS data from 270 liver cancers to the PCAWG (10% of the total), making us the most productive group within the ICGC. In 2018, as a PCAWG-15 working group, RIKEN and IMUT analyzed the immuno-genomic landscape from PCAWG data, including mutations in HLAs and immune suppressor genes, neo-antigen profiles and immune micro-environmental signatures, and observed that tumors acquired many types of immune escape mechanisms. As a PCAWG-10 working group, RIKEN also analyzed PCAWG WGS data for genome-wide somatic mutations in microsatellite or repeat regions, one of the most difficult regions to analyze by NGS, and defined microsatellite instability (MSI) cancers at a genome-wide level. They also identified highly mutated microsatellites that can be used to detect MSI cancers with high sensitivity.
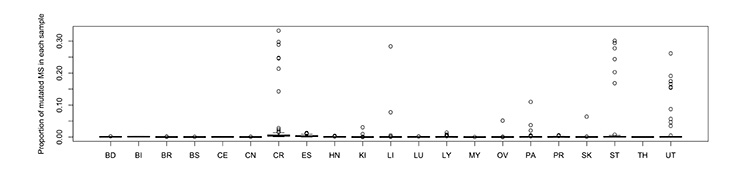
Figure1: Mutation rate of 7M microsatellite regions in each type of cancer.
CR (colorectal), ST (stomach) and UT (Uterine) cancers showed a higher mutation rate (>0.3% of 7M) in microsatellite regions, indicating microsatellite instability (MSI).