Genome Immunobiology RIKEN Hakubi Research Team
1. Immunobiology of Endogenous Viral Elements in Mammals
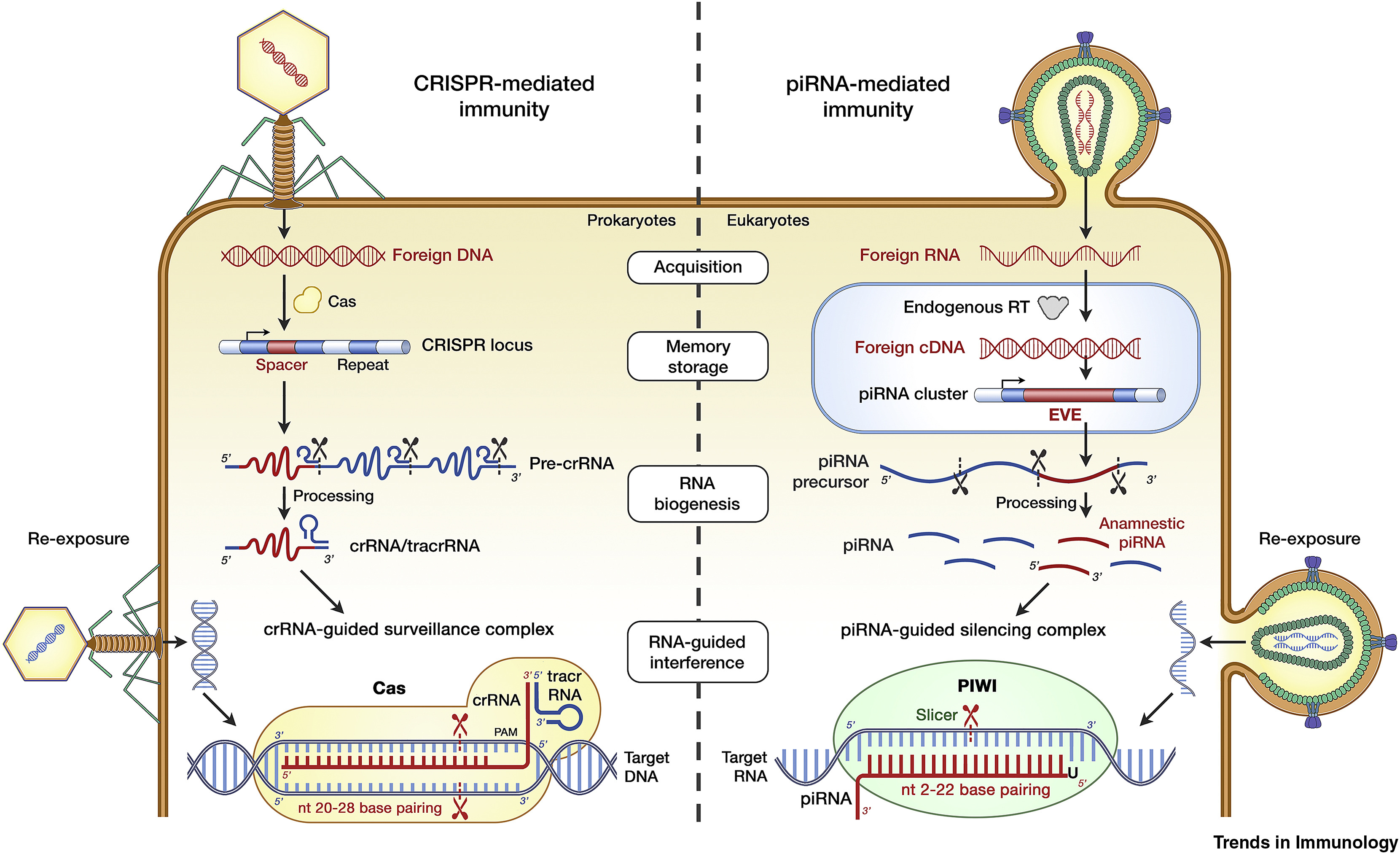
We are testing the hypothesis that small RNAs generated from virus-derived sequences called Endogenous Bornavirus-Like Nucleoprotein Elements (EBLNs) influence bornavirus infection (Parrish et al., RNA 2015). In this project we are using mice genetically engineered to lack EBLNs, to contain novel EBLN-like sequences, or to lack specific small RNA pathways. We are using bioinformatic approaches to test another prediction of this model: that other sequences in piRNA clusters may have evolved by horizontal gene transfer from viruses (Ophinni et al., Trends in Immunology 2019; Kojima et al., PNAS 2021).
2. Endogenization and excision of human herpesvirus 6 (HHV-6)
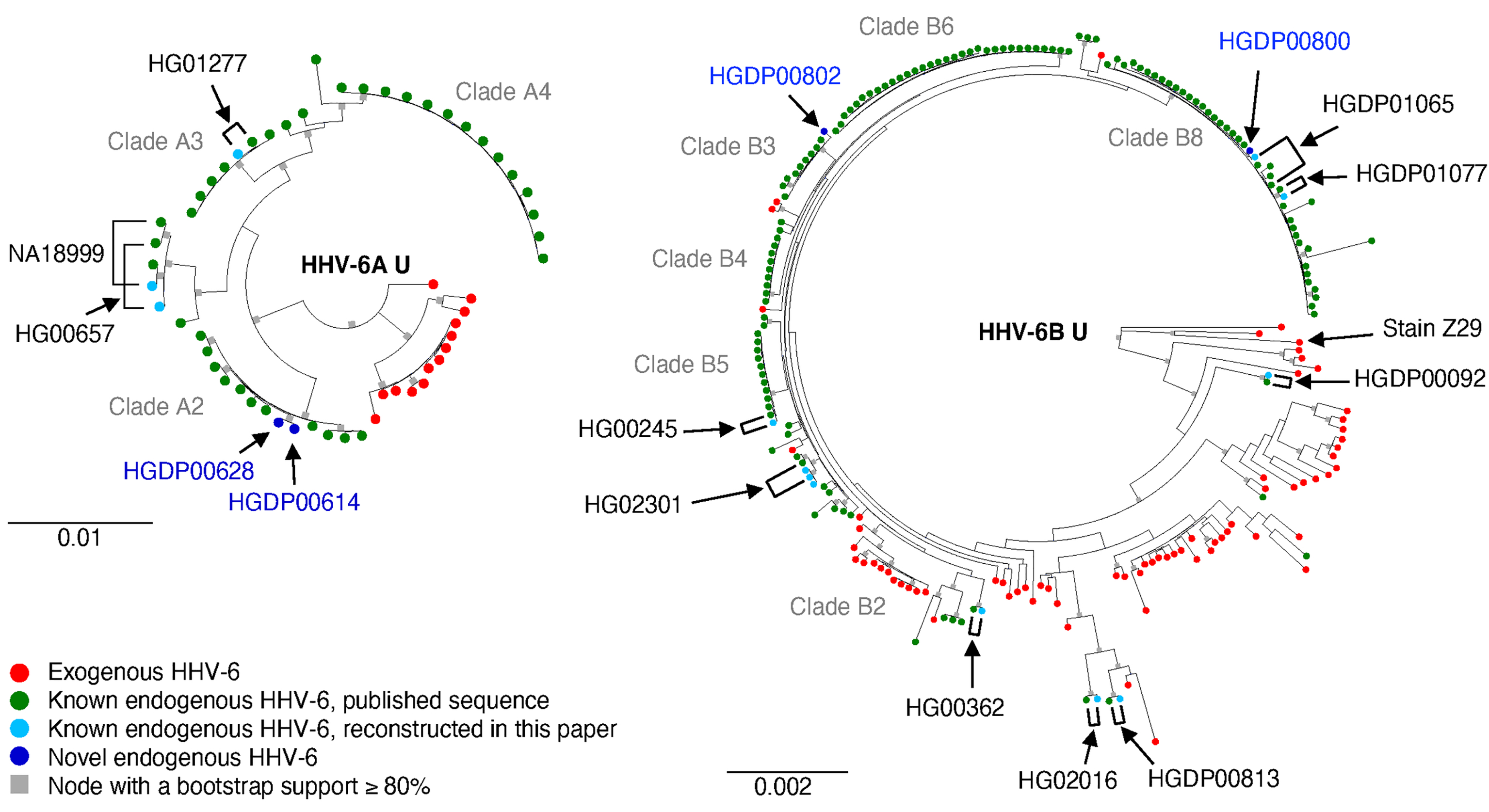
About 1% of people inherit sequences derived from HHV-6 in one of their chromosomes' telomeres; this is called endogenous HHV-6 or chromosomally-integrated HHV-6. HHV-6 is thought to be the most recently-acquired endogenous virus in human genomes (Aswad et al., Molecular Biology and Evolution 2020; Liu et al., PLoS Genetics 2020). Endogenization of HHV-6 may have some negative consequences on human health, including adverse pregnancy outcomes. We are using biobank-scale data to study the phenotypes, including potentially protective immune responses, associated with endogenous HHV-6 (Kojima et al., PLoS Genetics 2021). We also found evidence that HHV-6 has been excised from some human chromosomes via recombination (Liu et al., PLoS Genetics 2020). We are studying whether this excision can result in production of infectious virus from a latent endogenous reservoir; this would have several implications relevant to medicine, including organ and tissue transplantation.
3. Influence of MGEs on host genotype and phenotype
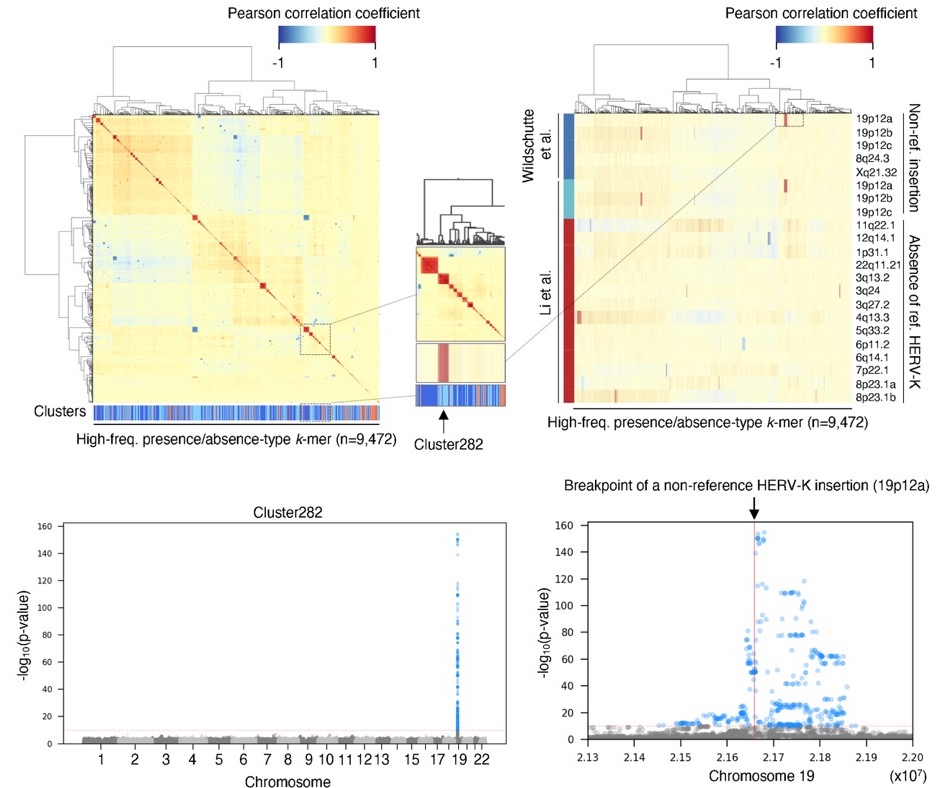
Viruses are the mobile genetic elements (MGEs) best recongized for causing variation in human phenotypes - most strikingly, diseases in people infected by them. But other MGEs, notably transposons, may also control variation in host phenotypes. MGE-derived sequences make up almost half of our genome, but their individual-to-individual variation is not yet exhaustively catalogued, nor are the mechanisms by which MGEs can influence genome, cell, and tissue function. We are coupling statistical genetic and bioinformatic approaches with long-read sequencing and genome editing to study how MGEs influence phenotypes across biological hierarchies (e.g. Sakashita et al., Nature Structural and Molecular Biology 2020; Ito et al., bioRxiv 2020; Kojima et al., PLoS Genetics 2021).
4. Retrobiology
Reverse transcription can stabilize the information content of an RNA molecule into a readily-heritable DNA form. Genome sequences acquired via reverse transcription are involved in diverse cellular and organismal processes (Parrish and Tomonaga, Current Opinion in Microbiology 2016), including some as complex as synaptic communication (Parrish and Tomonaga, Cell 2018). Working within the 1000 Genomes Project structural variation working group, we found new examples of such sequences that are variably present in human populations, suggesting that the processes responsible for creating these sequences are ongoing in humans (Sudmant et al., Nature 2015). We are interested in characterizing the contribution of reverse transcribed sequences to mammalian genomes and understanding the functions these sequences enable.
Images used under Creative Commons Attribution-Non-Commercial 4.0 International License (CC-BY-NC).