Projects
Integration of genetic information and immune functions
Recently, many disease susceptibility variants have been identified by genome wide association study (GWAS). Germline genetic variations provide us with evidence into the causal relationship of an observed phenomenon and its pathogenesis. In this regard, the majority of GWAS risk variants have been found to function as an expression-quantitative trait locus (e-QTL), regulating the expression levels of genes. Therefore, using genomic information, qualitative and quantitative analyses of transcriptomes together with epigenomes, we will better understand the pathogenic components of immunocompetent cells in various immune-mediated diseases.
We are now using a system to obtain various subtypes of leukocytes from peripheral blood mononuclear cells (PBMC) from heathy individuals in order to obtain the utmost unbiased relationship between genotypes and gene expression. Cell separation is performed using fluorescence-activated cell sorting into about 20 different subsets. Cells are then analyzed in the steady state or under stimulated conditions, such as with combinations of cytokines and cell surface receptor agonists to capture the dynamic responses of gene regulation. As the first step, genotyping as well as RNA-seq are performed. Subsequently, we also perform epigenetic analyses, specifically focusing on enhancers and promoters, as well as long non-coding RNAs. We believe that cap analysis of gene expression (CAGE), assay for transposase-accessible chromatin using sequencing (ATAC-seq) and analysis of several histone marks for each subset will be powerful tools for identifying the causal relationship between genetic variation and gene expression.
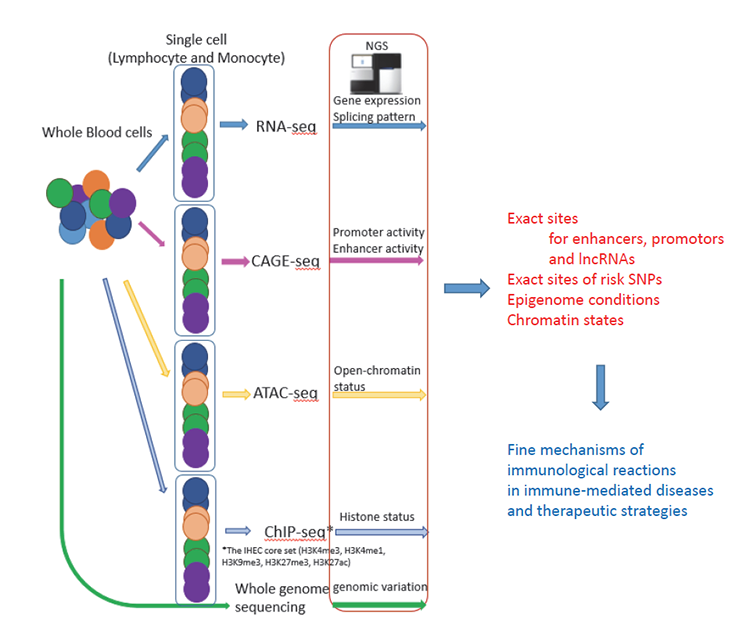
Figure: Integration of genetic information and immune functions: The eQTL project